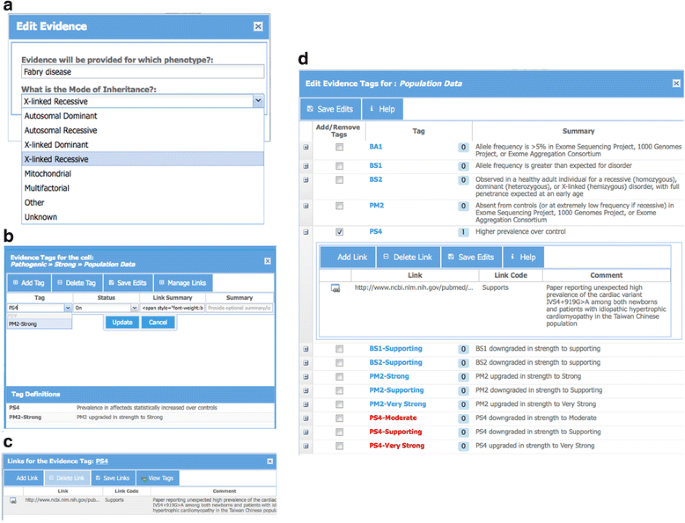
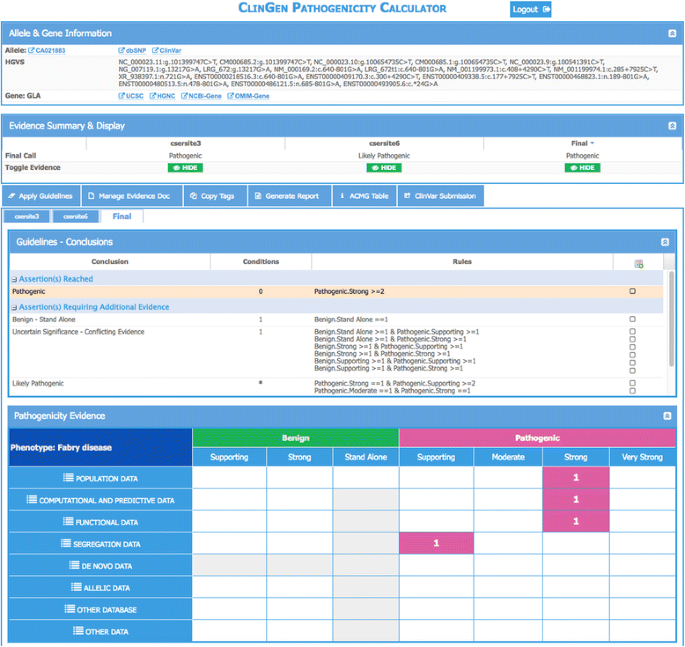
ClinGen Pathogenicity Calculator: a configurable system for assessing pathogenicity of genetic variants | Genome Medicine | Full Text
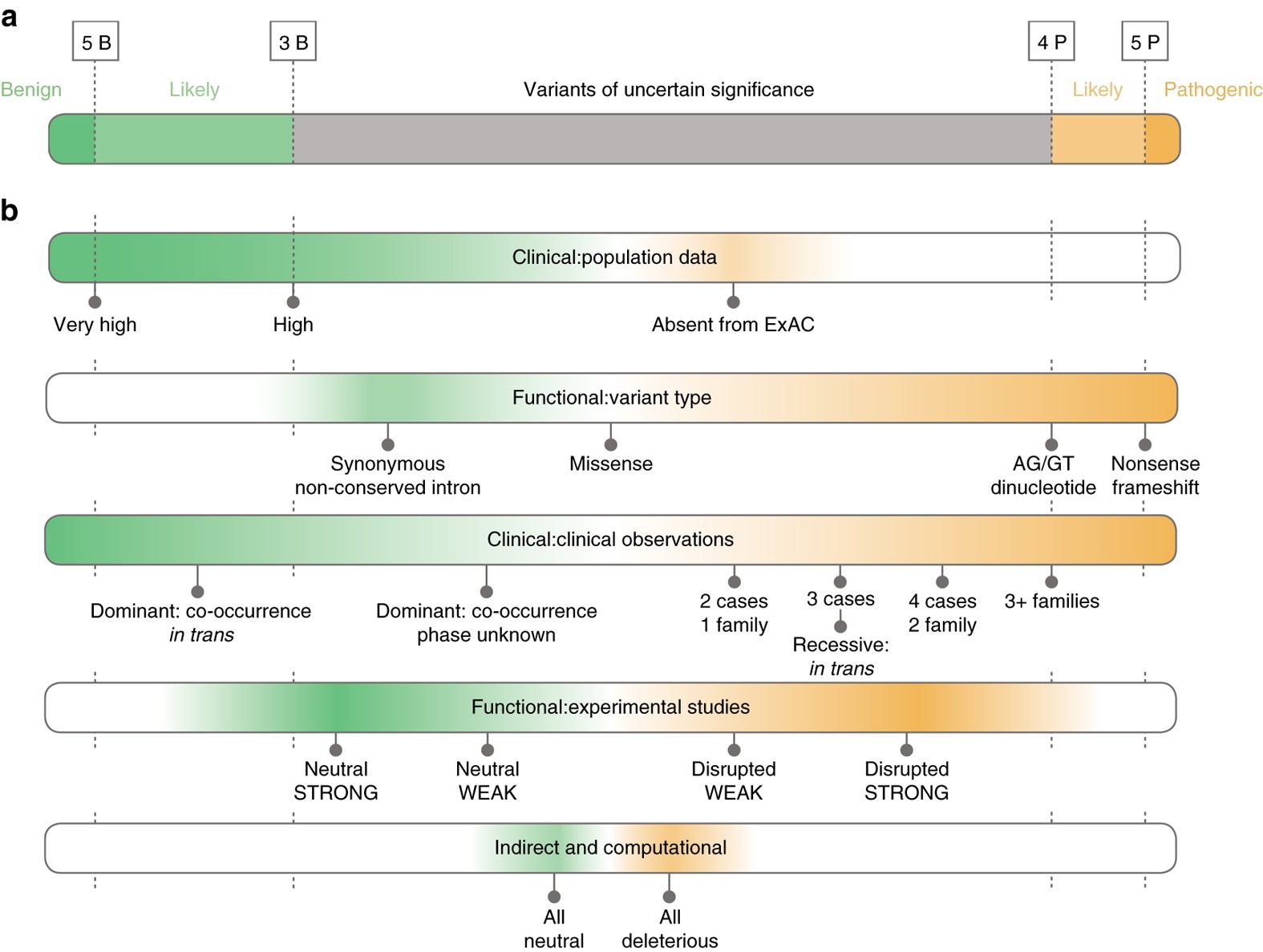
Sherloc: a comprehensive refinement of the ACMG–AMP variant classification criteria | Genetics in Medicine
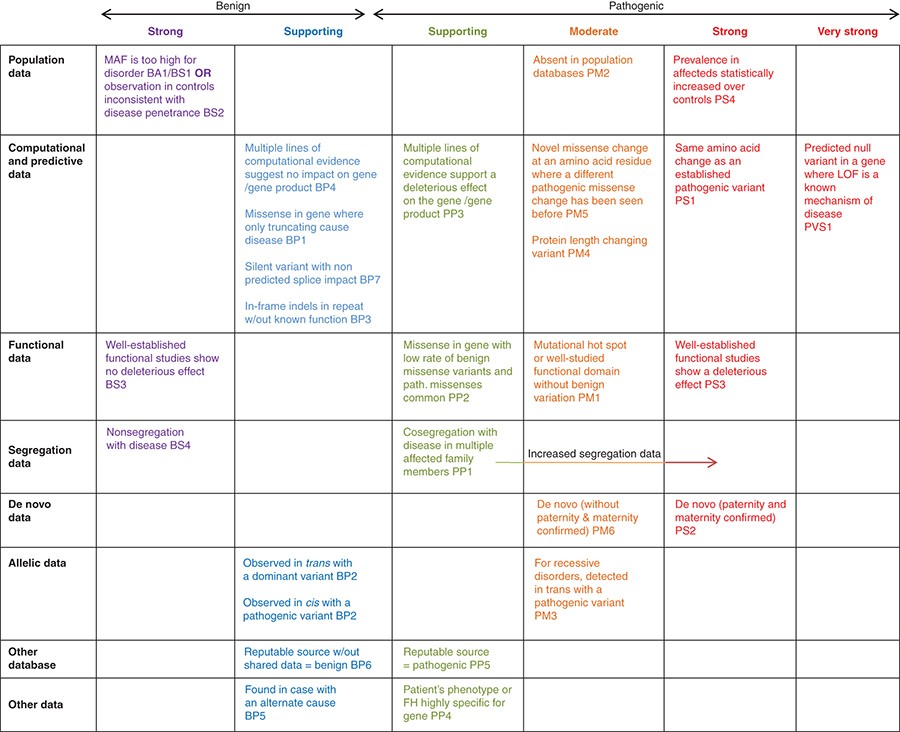
Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology | Genetics in Medicine

ACMG classification of NRAS G12S. (Top) ProteinPaint display of somatic... | Download Scientific Diagram

A Standardized DNA Variant Scoring System for Pathogenicity Assessments in Mendelian Disorders - Karbassi - 2016 - Human Mutation - Wiley Online Library

Consideration of Cosegregation in the Pathogenicity Classification of Genomic Variants - ScienceDirect

Cancer SIGVAR: A semi-automated interpretation tool for germline variants of hereditary cancer-related genes | bioRxiv

Classification and Reporting of Potentially Proarrhythmic Common Genetic Variation in Long QT Syndrome Genetic Testing | Circulation
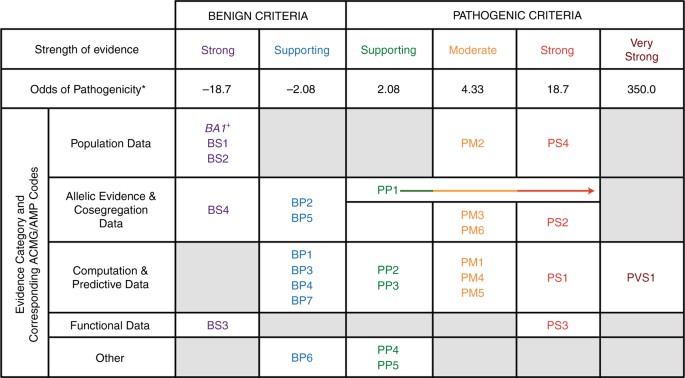
Navigating the nuances of clinical sequence variant interpretation in Mendelian disease | Genetics in Medicine
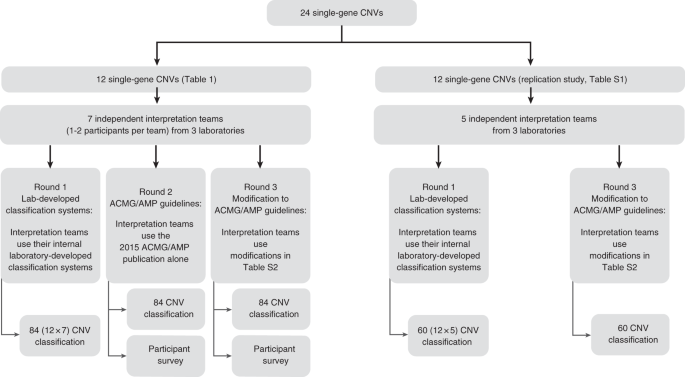
Adapting ACMG/AMP sequence variant classification guidelines for single-gene copy number variants | Genetics in Medicine
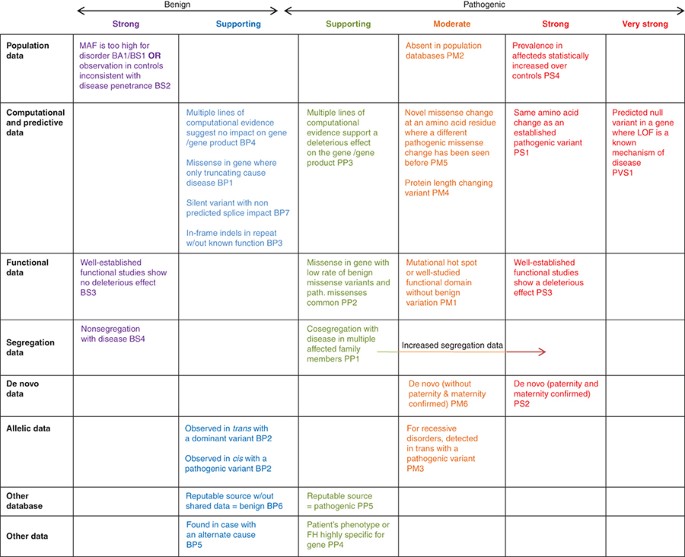
Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology | Genetics in Medicine

Proposed adaptation of ACMG/AMP guidelines for rule PM1, relating to... | Download Scientific Diagram

Navigating the nuances of clinical sequence variant interpretation in Mendelian disease - Genetics in Medicine

Current Tools, Databases, and Resources for Phenotype and Variant Analysis of Clinical Exome Sequencing - Advances in Molecular Pathology

InterVar: Clinical Interpretation of Genetic Variants by the 2015 ACMG-AMP Guidelines - ScienceDirect

Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer - The Journal of Molecular Diagnostics

AutoPVS1: An automatic classification tool for PVS1 interpretation of null variants - Xiang - 2020 - Human Mutation - Wiley Online Library






![PDF] ACGS Best Practice Guidelines for Variant Classification 2019 | Semantic Scholar PDF] ACGS Best Practice Guidelines for Variant Classification 2019 | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/86fa75f316823ef358a804fe1e45a98a85351bf9/4-Table1-1.png)